Tohban EOVSA Imaging Tutorial A-Z: Difference between revisions
| Line 4: | Line 4: | ||
If you are not using mobaxterm, directly SSH to Pipeline through your Linux or Mac computer and start tcsh to run CASA. | If you are not using mobaxterm, directly SSH to Pipeline through your Linux or Mac computer and start tcsh to run CASA. | ||
<pre style="background-color: #FCEBD9"> | |||
<pre style="background-color: #FCEBD9;overflow: auto;width: auto;"> | |||
from astropy.time import Time | from astropy.time import Time | ||
import os | import os | ||
Revision as of 02:44, 15 February 2022
Connection details to pipeline server
Step 1: Downloading raw data (IDB) on Pipeline machine
On CASA of the Pipeline machine, which has the complete EOVSA SQL database.
If you are not using mobaxterm, directly SSH to Pipeline through your Linux or Mac computer and start tcsh to run CASA.
from astropy.time import Time
import os
trange = Time(['2017-08-21 20:15:00', '2017-08-21 20:25:00']) ###Change accordingly###
#### (Optional) change output path, default current directory "./" #####
outpath = './msdata/' ###Change accordingly###
if not os.path.exists(outpath):
os.makedirs(outpath)
######################################################
msfiles = importeovsa(idbfiles=trange, ncpu=1, visprefix=outpath)
NOTE: The above step currently gives an error with importeovsa. This is because the data location was changed and the code doesn't know where to look for the IDB files. Sijie is looking into the problem. We will update the page when it is fixed. For now, please use the method below for all time ranges.
OR (for a time range longer than 10 minutes)
from suncasa.tasks import task_calibeovsa as calibeovsa
from suncasa.tasks import task_importeovsa as timporteovsa
from split_cli import split_cli as split
import dump_tsys as dt
from util import Time
import numpy as np
import os
from glob import glob
from eovsapy import util
trange = Time(['2017-08-21 20:15:00', '2017-08-21 20:35:00']) ###Change accordingly###
idbdir = util.get_idbdir(trange[0])
info = dt.rd_fdb(trange[0])
for k, v in info.iteritems():
info[k] = info[k][~(info[k] == '')]
sidx = np.where(
np.logical_and(info['SOURCEID'] == 'Sun', info['PROJECTID'] == 'NormalObserving') & np.logical_and(
info['ST_TS'].astype(np.float) >= trange[0].lv,
info['ST_TS'].astype(np.float) <= trange[
1].lv))
filelist = info['FILE'][sidx]
outpath = './msdata/' ###Change accordingly###
if not os.path.exists(outpath):
os.makedirs(outpath)
inpath = idbdir + '{}/'.format(trange[0].datetime.strftime("%Y%m%d"))
ncpu = 1
msfiles = timporteovsa.importeovsa(idbfiles=[inpath + ll for ll in filelist], ncpu=ncpu, timebin="0s", width=1,
visprefix=outpath,
nocreatms=False, doconcat=False,
modelms="", doscaling=False, keep_nsclms=False, udb_corr=True)
If you come across errors with calibeovsa, add following lines to your ~/.casa/init.py file.
import sys
sys.path.append('/common/python')
sys.path.append('/common/python/packages/pipeline_casa')
execfile('/common/python/suncasa/tasks/mytasks.py')
Step 2: Concatenate all the 10 mins data, if there are any
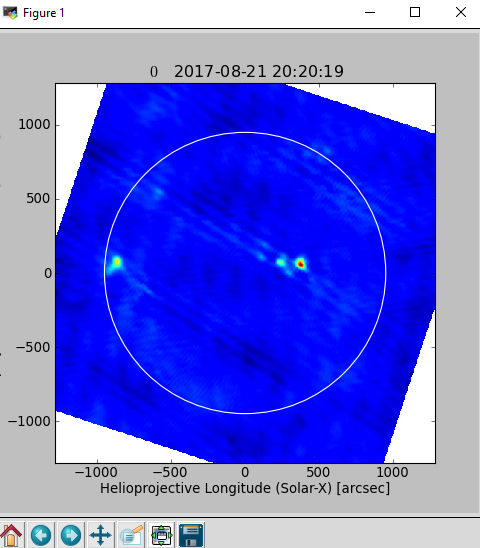
Follow this step if there are more than one .ms files, if not run step 3 directly. If doimage=True, a quicklook image will be produced (by integrating over the entire time) as shown below.
# This is to set the path/name for the concatenated files concatvis = os.path.basename(msfiles[0])[:11] + '_concat_cal.ms' vis = calibeovsa.calibeovsa(msfiles, doconcat=True, concatvis=concatvis, caltype=['refpha','phacal'], doimage=True)
Step 3: Calibration
This will calibrate the input visibility, write out calibration tables under /data1/eovsa/caltable/, and apply the calibration.
vis = calibeovsa.calibeovsa('IDB20170821202020.ms', caltype=['refpha','phacal'], doimage=True) ###Change the vis filename accordingly###
# Append '_cal' to the ms filename and split the corrected column to the new caled ms
vis_str = str(' '.join(vis))
caled_vis=vis_str.replace('.ms','_cal.ms')
split(vis=' '.join(vis),outputvis=caled_vis,datacolumn='corrected',timerange='',correlation='XX')
One needs to transfer the created caled ms data files (xxx_concat_cal.ms or xxx_cal.ms) from pipeline to inti server, which has all the casatasks installed, in order to run the rest of the imaging steps.
Connection details to Inti server
For Windows, on Mobaxterm,
- With your NJIT VPN connected, connect to one of the afsconnect servers (for example, afsaccess3.njit.edu) using your UCID and password.
- ssh -X UCID@inti.hpcnet.campus.njit.edu
To avoid typing the full inti address each time you attempt for ssh, you may wish to add the following lines with your username to C:\Program Files\Git\etc\ssh\ssh_config on Windows and /.ssh/config on Mac and Linux machines.
Host inti Hostname inti.hpcnet.campus.njit.edu User USERNAME
To have sufficient disk space with EOVSA data analysis over Inti, use your dedicated directory YOURDIRECTORY at the location given below. If you do not have a directory, Please take help from Sijie in creating one.
cd /inti/data/users/YOURDIRECTORY
For Linux or Mac machine, ssh -X UCID@inti.hpcnet.campus.njit.edu
Transferring data files between servers
For directly transferring your calibrated .ms data between the Pipeline and Inti servers, follow the below given steps.
1. Log into Inti using your username. ssh -X USERNAME@inti.hpcnet.campus.njit.edu Then create a tunnel into Pipeline from Inti. ssh -L 8888:pipeline.solar.pvt:22 guest@ovsa.njit.edu 2. Log into Inti again from a new terminal. Change to your working directory and give this command to copy your data on Pipeline. scp -v -C -r -P 8888 USERNAMEofPipeline@localhost:PATHofMSDATA/MSfilename ./ Eg: scp -v -C -r -P 8888 shaheda@localhost:/data1/shaheda/IDB20220118173922_cal.ms ./ where, MSfilename, PATHofMSDATA are your .ms data and its path on Pipeline machine.
Alternatively, one can follow the below procedure to do the transfer.
For Windows, on mobaxterm, drag and drop the ms file to your local machine or use scp command as given below.
scp -r -C userid@pipeline:/your/folder/msfile /localmachine/destination/folder/
On mobaxterm and on your local terminal, use the following command to finally copy the ms file to Inti.
scp -r -C /localmachine/destination/folder/msfile UCID@inti.hpcnet.campus.njit.edu:/inti/data/users/YOURDIRECTORY
For Mac and Linux, SCP can be used in the same way. Add the following lines to your SSH config file, to bypass ovsa.njit.edu from copying. vi ~/.ssh/config
Host ovsa
HostName ovsa.njit.edu
User guest
Host pipeline
Hostname pipeline.solar.pvt
User userid
ProxyCommand ssh -W %h:%p guest@ovsa.njit.edu
Software details on the servers
On Inti, when logging in for the first time, please add the following lines to your accounts ~/.bashrc file.
>>vi ~/.bashrc Insert the text given below and save it.
#### setting start ####
if [ $HOSTNAME == "baozi.hpcnet.campus.njit.edu" ]; then
source /srg/.setenv_baozi
fi
if [ $HOSTNAME == "inti.hpcnet.campus.njit.edu" ]; then
source /inti/.setenv_inti
fi
if [ $HOSTNAME == "guko.resource.campus.njit.edu" ]; then
source /data/data/.setenv_guko
fi
#### setting end ####
Both CASA 5 and 6 are available on Inti.
Enter the bash environment on inti, and load the desired casa environment. To load CASA 5, enter bash environment by giving >>bash
>> loadcasa5 >> suncasa #This should load the software making you ready for the analysis
Or to load CASA 6, enter bash environment by giving >>bash
>> loadcasa6 >> ipython #This should load the software making you ready for the analysis
Here, for example, to use clean, first start ipython as given above, then type in >>from casatasks import tclean
Step 4: Self-calibration on Inti server
https://github.com/binchensun/casa-eovsa/blob/master/slfcal_example.py
from suncasa.utils import helioimage2fits as hf
import os
import numpy as np
import pickle
from matplotlib import gridspec as gridspec
from sunpy import map as smap
from matplotlib import pyplot as plt
from split_cli import split_cli as split
import time
# =========== task handlers =============
dofullsun = 1 # initial full-sun imaging ###Change accordingly###
domasks=1 # get masks ###Change accordingly###
doslfcal=1 # main cycle of doing selfcalibration ###Change accordingly###
doapply=1 # apply the results ###Change accordingly###
doclean_slfcaled=1 # perform clean for self-calibrated data ###Change accordingly###
# ============ declaring the working directories ============
workdir = os.getcwd()+'/' #main working directory. Using current directory in this example
slfcaldir = workdir+'slfcal/' #place to put all selfcalibration products
imagedir = slfcaldir+'images/' #place to put all selfcalibration images
maskdir = slfcaldir+'masks/' #place to put clean masks
imagedir_slfcaled = slfcaldir+'images_slfcaled/' #place to put final self-calibrated images
caltbdir = slfcaldir+'caltbs/' # place to put calibration tables
# make these directories if they do not already exist
dirs = [workdir, slfcaldir, imagedir, maskdir, imagedir_slfcaled, caltbdir]
for d in dirs:
if not os.path.exists(d):
os.makedirs(d)
# ============ Split a short time for self-calibration ===========
# input visibility
ms_in = workdir + 'IDB20170821202020_cal.ms' ###Change the initial calibrated (through calibeovsa) vis accordingly###
# output, selfcaled, visibility
ms_slfcaled = workdir + os.path.basename(ms_in).replace('cal','slfcaled')
# intermediate small visibility for selfcalbration
# selected time range for generating self-calibration solutions
trange='2017/08/21/20:21:10~2017/08/21/20:21:30' ###Change accordingly###
slfcalms = slfcaldir+'slfcalms.XX.slfcal'
slfcaledms = slfcaldir+'slfcalms.XX.slfcaled'
if not os.path.exists(slfcalms):
split(vis=ms_in,outputvis=slfcalms,datacolumn='data',timerange=trange,correlation='XX')
# ============ Prior definitions for spectral windows, antennas, pixel numbers =========
spws=[str(s+1) for s in range(30)]
antennas='0~12'
npix=512
nround=3 #number of slfcal cycles
# =========== Step 1, doing a full-Sun image to find out phasecenter and appropriate field of view =========
if dofullsun:
#initial mfs clean to find out the image phase center
im_init='fullsun_init'
os.system('rm -rf '+im_init+'*')
tclean(vis=slfcalms,
antenna='0~12',
imagename=im_init,
spw='1~15',
specmode='mfs',
timerange=trange,
imsize=[npix],
cell=['5arcsec'],
niter=1000,
gain=0.05,
stokes='I',
restoringbeam=['30arcsec'],
interactive=False,
pbcor=True)
hf.imreg(vis=slfcalms,imagefile=im_init+'.image.pbcor',fitsfile=im_init+'.fits',
timerange=trange,usephacenter=False,verbose=True)
clnjunks = ['.flux', '.mask', '.model', '.psf', '.residual','.sumwt','.pb','.image']
for clnjunk in clnjunks:
if os.path.exists(im_init + clnjunk):
os.system('rm -rf '+im_init + clnjunk)
from sunpy import map as smap
from matplotlib import pyplot as plt
fig = plt.figure(figsize=(6,6))
ax = fig.add_subplot(111)
eomap=smap.Map(im_init+'.fits')
#eomap.data=eomap.data.reshape((npix,npix))
eomap.plot_settings['cmap'] = plt.get_cmap('jet')
eomap.plot(axes = ax)
eomap.draw_limb()
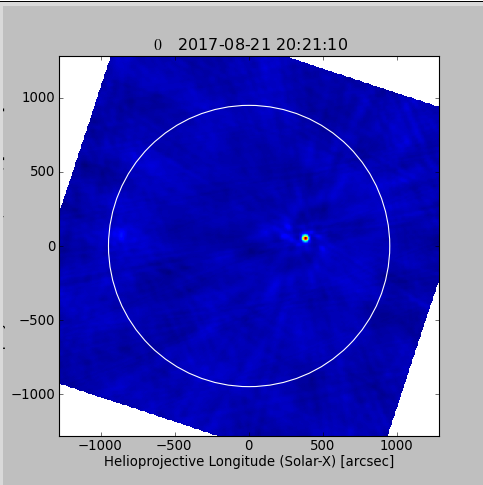
plt.show()
viewer(im_init+'.image.pbcor')
# parameters specific to the event (found from step 1)
phasecenter='J2000 10h02m59 11d58m07' ###Change accordingly###
xran=[280,480] ###Change accordingly###
yran=[-50,150] ###Change accordingly###
# =========== Step 2 (optional), generate masks =========
# if skipped, will not use any masks
if domasks:
clearcal(slfcalms)
delmod(slfcalms)
antennas='0~12'
pol='XX'
imgprefix=maskdir+'slf_t0'
# step 1: set up the clean masks
img_init=imgprefix+'_init_ar_'
os.system('rm -rf '+img_init+'*')
#spwrans_mask=['1~5','6~12','13~20','21~30']
spwrans_mask=['1~12']
#convert to a list of spws
spwrans_mask_list = [[str(i) for i in (np.arange(int(m.split('~')[0]),int(m.split('~')[1])))] for m in spwrans_mask]
masks=[]
imnames=[]
for spwran in spwrans_mask:
imname=img_init+spwran.replace('~','-')
try:
tclean(vis=slfcalms,
antenna='0~12',
imagename=imname,
spw=spwran,
specmode='mfs',
timerange=trange,
imsize=[npix],
cell=['2arcsec'],
niter=1000,
gain=0.05,
stokes='XX',
restoringbeam=['20arcsec'],
phasecenter=phasecenter,
weighting='briggs',
robust=1.0,
interactive=True,
datacolumn='data',
pbcor=True,
savemodel='modelcolumn')
imnames.append(imname+'.image')
masks.append(imname+'.mask')
clnjunks = ['.flux', '.model', '.psf', '.residual']
for clnjunk in clnjunks:
if os.path.exists(imname + clnjunk):
os.system('rm -rf '+ imname + clnjunk)
except:
print('error in cleaning spw: '+spwran)
pickle.dump(masks,open(slfcaldir+'masks.p','wb'))
# =========== Step 3, main step of selfcalibration =========
if doslfcal:
if os.path.exists(slfcaldir+'masks.p'):
masks=pickle.load(open(slfcaldir+'masks.p','rb'))
if not os.path.exists(slfcaldir+'masks.p'):
print 'masks do not exist. Use default mask'
masks=[]
os.system('rm -rf '+imagedir+'*')
os.system('rm -rf '+caltbdir+'*')
#first step: make a mock caltable for the entire database
print('Processing ' + trange)
slftbs=[]
calprefix=caltbdir+'slf'
imgprefix=imagedir+'slf'
tb.open(slfcalms+'/SPECTRAL_WINDOW')
reffreqs=tb.getcol('REF_FREQUENCY')
bdwds=tb.getcol('TOTAL_BANDWIDTH')
cfreqs=reffreqs+bdwds/2.
tb.close()
# starting beam size at 3.4 GHz in arcsec
sbeam=40.
strtmp=[m.replace(':','') for m in trange.split('~')]
timestr='t'+strtmp[0]+'-'+strtmp[1]
refantenna='0'
# number of iterations for each round
niters=[100, 300, 500]
# roburst value for weighting the baselines
robusts=[1.0, 0.5, 0.0]
# apply calibration tables? Set to true for most cases
doapplycal=[1,1,1]
# modes for calibration, 'p' for phase-only, 'a' for amplitude only, 'ap' for both
calmodes=['p','p','a']
# setting uvranges for model image (optional, not used here)
uvranges=['','','']
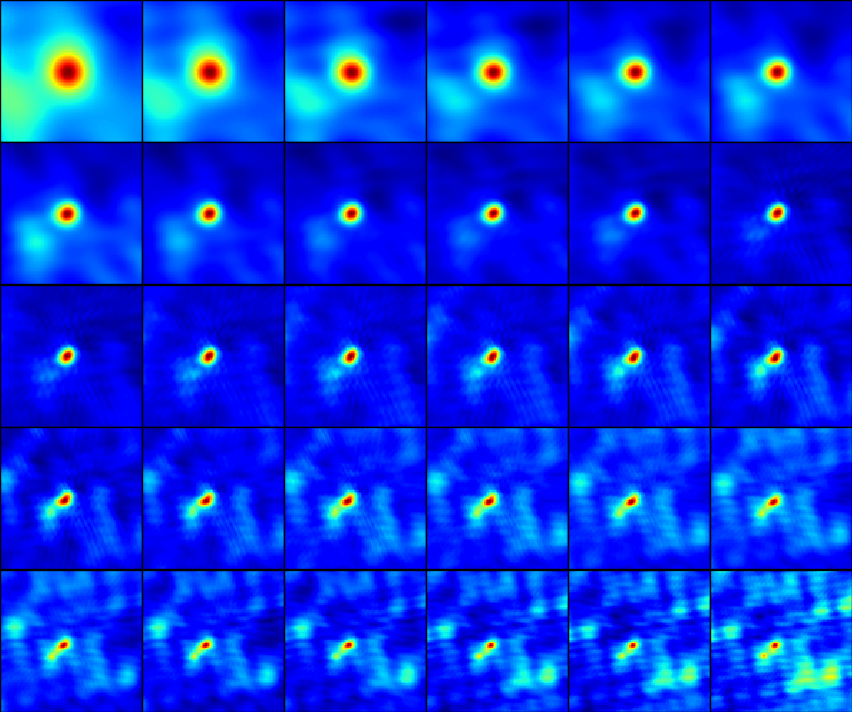
for n in range(nround):
slfcal_tb_g= calprefix+'.G'+str(n)
fig = plt.figure(figsize=(8.4,7.))
gs = gridspec.GridSpec(5, 6)
for s,sp in enumerate(spws):
print 'processing spw: '+sp
cfreq=cfreqs[int(sp)]
# setting restoring beam size (not very useful for selfcal anyway, but just to see the results)
bm=max(sbeam*cfreqs[1]/cfreq, 6.)
slfcal_img = imgprefix+'.spw'+sp.zfill(2)+'.slfcal'+str(n)
# only the first round uses nearby spws for getting initial model
if n == 0:
spbg=max(int(sp)-2,1)
sped=min(int(sp)+2,30)
spwran=str(spbg)+'~'+str(sped)
print('using spw {0:s} as model'.format(spwran))
if 'spwrans_mask' in vars():
for m, spwran_mask in enumerate(spwrans_mask):
if sp in spwran_mask:
mask = masks[m]
print('using mask {0:s}'.format(mask))
findmask = True
if not findmask:
print('mask not found. Do use any masks')
else:
spwran = sp
if 'spwrans_mask' in vars():
for m, spwran_mask in enumerate(spwrans_mask):
if sp in spwran_mask:
mask = masks[m]
print 'using mask {0:s}'.format(mask)
findmask = True
if not findmask:
print('mask not found. Do use any masks')
try:
tclean(vis=slfcalms,
antenna=antennas,
imagename=slfcal_img,
uvrange=uvranges[n],
spw=spwran,
specmode='mfs',
timerange=trange,
imsize=[npix],
cell=['2arcsec'],
niter=niters[n],
gain=0.05,
stokes='XX', #use pol XX image as the model
weighting='briggs',
robust=robusts[n],
phasecenter=phasecenter,
mask=mask,
restoringbeam=[str(bm)+'arcsec'],
pbcor=False,
interactive=False,
savemodel='modelcolumn')
if os.path.exists(slfcal_img+'.image'):
fitsfile=slfcal_img+'.fits'
hf.imreg(vis=slfcalms,imagefile=slfcal_img+'.image',fitsfile=fitsfile,
timerange=trange,usephacenter=False,toTb=True,verbose=False,overwrite=True)
clnjunks = ['.mask','.flux', '.model', '.psf', '.residual', '.image','.pb','.image.pbcor','.sumwt']
for clnjunk in clnjunks:
if os.path.exists(slfcal_img + clnjunk):
os.system('rm -rf '+ slfcal_img + clnjunk)
ax = fig.add_subplot(gs[s])
eomap=smap.Map(fitsfile)
eomap.plot_settings['cmap'] = plt.get_cmap('jet')
eomap.plot(axes = ax)
eomap.draw_limb()
#eomap.draw_grid()
ax.set_title(' ')
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
ax.set_xlim(xran)
ax.set_ylim(yran)
os.system('rm -f '+ fitsfile)
except:
print 'error in cleaning spw: '+sp
print 'using nearby spws for initial model'
sp_e=int(sp)+2
sp_i=int(sp)-2
if sp_i < 1:
sp_i = 1
if sp_e > 30:
sp_e = 30
sp_=str(sp_i)+'~'+str(sp_e)
try:
tget(tclean)
spw=sp_
print('using spw {0:s} as model'.format(sp_))
tclean()
except:
print 'still not successful. abort...'
break
gaincal(vis=slfcalms, refant=refantenna,antenna=antennas,caltable=slfcal_tb_g,spw=sp, uvrange='',\
gaintable=[],selectdata=True,timerange=trange,solint='inf',gaintype='G',calmode=calmodes[n],\
combine='',minblperant=4,minsnr=2,append=True)
if not os.path.exists(slfcal_tb_g):
print 'No solution found in spw: '+sp
figname=imagedir+'slf_t0_n{:d}.png'.format(n)
plt.subplots_adjust(left=0, bottom=0, right=1, top=1, wspace=0, hspace=0)
plt.savefig(figname)
time.sleep(10)
plt.close()
if os.path.exists(slfcal_tb_g):
slftbs.append(slfcal_tb_g)
slftb=[slfcal_tb_g]
os.chdir(slfcaldir)
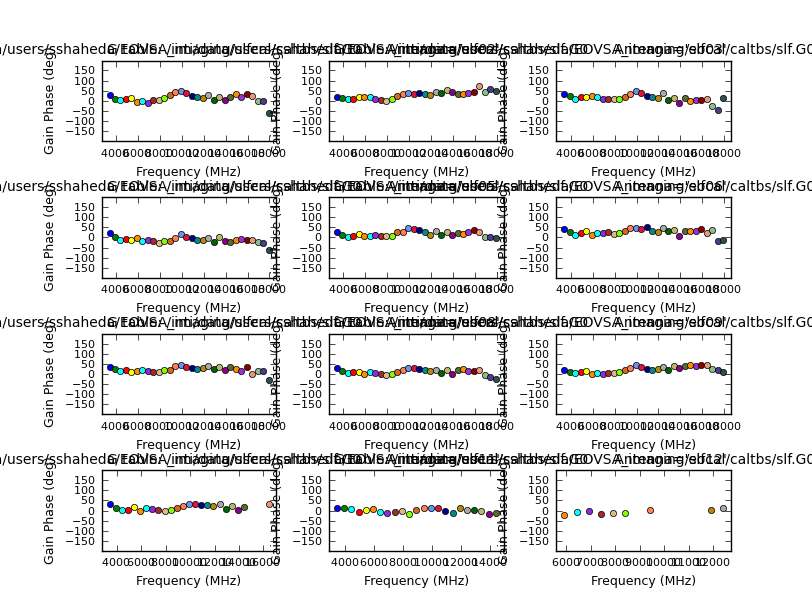
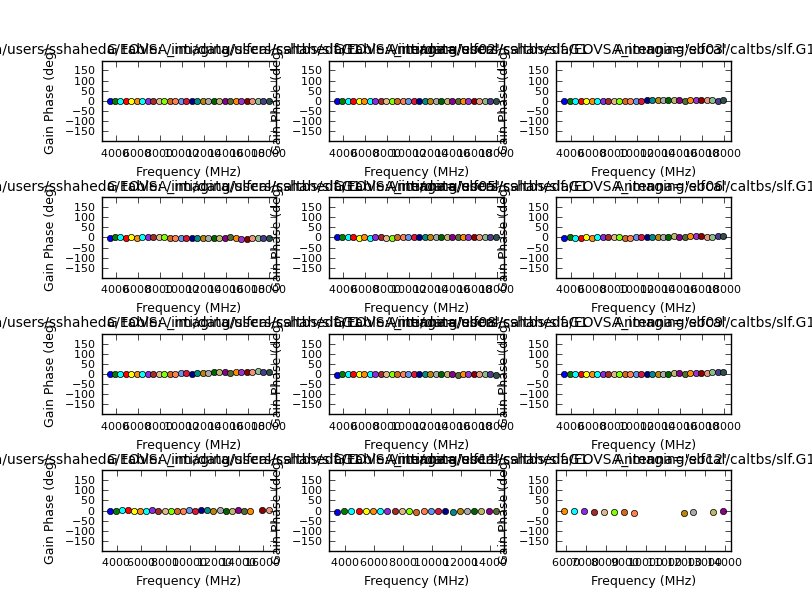
if calmodes[n] == 'p':
plotcal(caltable=slfcal_tb_g,antenna='1~12',xaxis='freq',yaxis='phase',\
subplot=431,plotrange=[-1,-1,-180,180],iteration='antenna',figfile=slfcal_tb_g+'.png',showgui=False)
if calmodes[n] == 'a':
plotcal(caltable=slfcal_tb_g,antenna='1~12',xaxis='freq',yaxis='amp',\
subplot=431,plotrange=[-1,-1,0,2.],iteration='antenna',figfile=slfcal_tb_g+'.png',showgui=False)
os.chdir(workdir)
if doapplycal[n]:
clearcal(slfcalms)
delmod(slfcalms)
applycal(vis=slfcalms,gaintable=slftb,spw=','.join(spws),selectdata=True,\
antenna=antennas,interp='nearest',flagbackup=False,applymode='calonly',calwt=False)
if n < nround-1:
prompt=raw_input('Continuing to selfcal?')
#prompt='y'
if prompt.lower() == 'n':
if os.path.exists(slfcaledms):
os.system('rm -rf '+slfcaledms)
split(slfcalms,slfcaledms,datacolumn='corrected')
print 'Final calibrated ms is {0:s}'.format(slfcaledms)
break
if prompt.lower() == 'y':
slfcalms_=slfcalms+str(n)
if os.path.exists(slfcalms_):
os.system('rm -rf '+slfcalms_)
split(slfcalms,slfcalms_,datacolumn='corrected')
slfcalms=slfcalms_
else:
if os.path.exists(slfcaledms):
os.system('rm -rf '+slfcaledms)
split(slfcalms,slfcaledms,datacolumn='corrected')
print 'Final calibrated ms is {0:s}'.format(slfcaledms)
# =========== Step 4: Apply self-calibration tables =========
if doapply:
import glob
os.chdir(workdir)
clearcal(ms_in)
clearcal(slfcalms)
applycal(vis=slfcalms,gaintable=slftbs,spw=','.join(spws),selectdata=True,\
antenna=antennas,interp='linear',flagbackup=False,applymode='calonly',calwt=False)
applycal(vis=ms_in,gaintable=slftbs,spw=','.join(spws),selectdata=True,\
antenna=antennas,interp='linear',flagbackup=False,applymode='calonly',calwt=False)
if os.path.exists(ms_slfcaled):
os.system('rm -rf '+ms_slfcaled)
split(ms_in, ms_slfcaled,datacolumn='corrected')
# =========== Step 5: Generate final self-calibrated images (optional) =========
if doclean_slfcaled:
import glob
pol='XX'
print('Processing ' + trange)
img_final=imagedir_slfcaled+'/slf_final_{0}_t0'.format(pol)
vis = ms_slfcaled
spws=[str(s+1) for s in range(30)]
tb.open(vis+'/SPECTRAL_WINDOW')
reffreqs=tb.getcol('REF_FREQUENCY')
bdwds=tb.getcol('TOTAL_BANDWIDTH')
cfreqs=reffreqs+bdwds/2.
tb.close()
sbeam=30.
from matplotlib import gridspec as gridspec
from sunpy import map as smap
from matplotlib import pyplot as plt
fitsfiles=[]
for s,sp in enumerate(spws):
cfreq=cfreqs[int(sp)]
bm=max(sbeam*cfreqs[1]/cfreq,4.)
imname=img_final+'_s'+sp.zfill(2)
fitsfile=imname+'.fits'
if not os.path.exists(fitsfile):
print 'cleaning spw {0:s} with beam size {1:.1f}"'.format(sp,bm)
try:
tclean(vis=vis,
antenna=antennas,
imagename=imname,
spw=sp,
specmode='mfs',
timerange=trange,
imsize=[npix],
cell=['1arcsec'],
niter=1000,
gain=0.05,
stokes=pol,
weighting='briggs',
robust=2.0,
restoringbeam=[str(bm)+'arcsec'],
phasecenter=phasecenter,
mask='',
pbcor=True,
interactive=False)
except:
print 'cleaning spw '+sp+' unsuccessful. Proceed to next spw'
continue
if os.path.exists(imname+'.image.pbcor'):
imn = imname+'.image.pbcor'
hf.imreg(vis=vis,imagefile=imn,fitsfile=fitsfile,
timerange=trange,usephacenter=False,toTb=True,verbose=False)
fitsfiles.append(fitsfile)
junks=['.flux','.model','.psf','.residual','.mask','.image','.pb','.image.pbcor','.sumwt']
for junk in junks:
if os.path.exists(imname+junk):
os.system('rm -rf '+imname+junk)
else:
print('fits file '+fitsfile+' already exists, skip clean...')
fitsfiles.append(fitsfile)
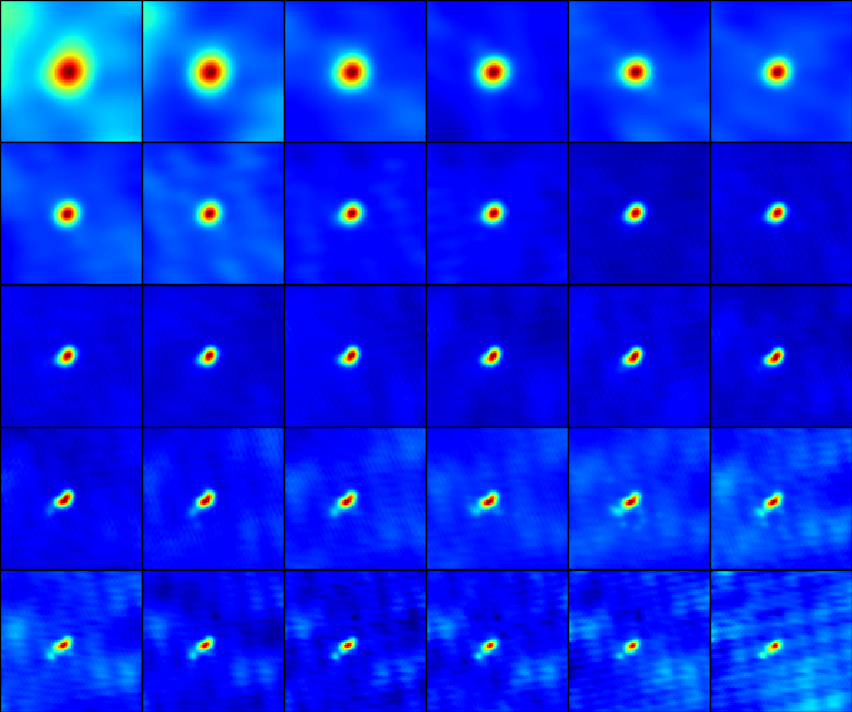
fig = plt.figure(figsize=(8.4,7.))
gs = gridspec.GridSpec(5, 6)
for s,sp in enumerate(spws):
cfreq=cfreqs[int(sp)]
ax = fig.add_subplot(gs[s])
eomap=smap.Map(fitsfiles[s])
eomap.plot_settings['cmap'] = plt.get_cmap('jet')
eomap.plot(axes = ax)
eomap.draw_limb()
ax.set_title(' ')
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
ax.set_xlim(xran)
ax.set_ylim(yran)
plt.text(0.98,0.85,'{0:.1f} GHz'.format(cfreq/1e9),transform=ax.transAxes,ha='right',color='w',fontweight='bold')
plt.subplots_adjust(left=0, bottom=0, right=1, top=1, wspace=0, hspace=0)
plt.show()
The .fits files of the self calibrated images at each frequency for the given time are saved at /slfcal/images_slfcaled in your working directory.
Step 5: Quick-look imaging
For further analysis of the event, follow the link given below. https://colab.research.google.com/drive/1lSLLxgOG6b8kgu9Sk6kSKvrViyubnXG6?usp=sharing#scrollTo=xbXyyLmCFCGL
from suncasa.utils import qlookplot as ql ## (Optional) Supply the npz file of the dynamic spectrum from previous step.
## If not provided, the program will generate a new one from the visibility.
## set the time interval
from suncasa import dspec as ds
import time
visibility_data = 'IDB20170821202020_slfcaled.ms'
specfile = visibility_data + '.dspec.npz'
d = ds.Dspec(visibility_data, bl='4&9', specfile=specfile)
timerange = '19:02:00~19:02:10' ## select frequency range from 2.5 GHz to 3.5 GHz
spw = '2~5' ## select stokes XX
stokes = 'XX' ## turn off AIA image plotting, default is True
plotaia = False
ql.qlookplot(vis=visibility_data, specfile=specfile, timerange=timerange, spw=spw, \
stokes=stokes, plotaia=plotaia, restoringbeam=['60arcsec'], robust=0.5)